BONN, Germany/PHILADELPHIA, US: It has long been known that the gene TP63 can contribute to the development of cleft lip and palate, but the exact process has been unclear. A joint study by the universities of Bonn and Pennsylvania has now clarified how this mechanism works.
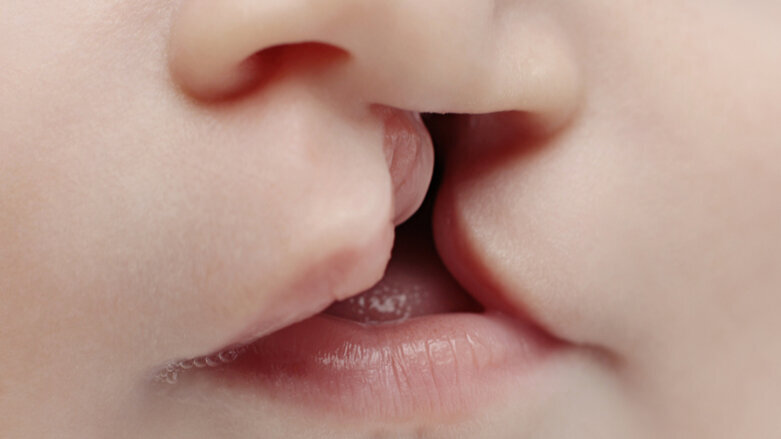
In Europe, about one in 500 babies are born with a cleft lip and palate. The cause is a developmental disorder in the first weeks of embryonic development. As a result, parts of the face or palate do not grow together as normal.
It has long been known that mutations of the TP63 gene can trigger particularly severe forms of this disorder. Those affected suffer not only from cleft formation in the facial area but also from malformations of the extremities and diseases of the skin, hair and teeth. Originally, it was assumed that TP63 only plays a role in the development of this syndromic form of cleft lip and palate.
“In the last two years, however, there has been increasing evidence that this is not the case,” said co-author Dr Kerstin Ludwig from the Institute of Human Genetics at the University Hospital of Bonn. “In our work, we were able to show for the first time that TP63 actually represents a link between the syndromic and the isolated form and how it interferes with facial development.”
Researchers at the University of Pennsylvania have succeeded in programming human cells to become facial cells in culture. This enabled the team to analyse the effect of TP63 on this particular cell type. This data was then combined by the Bonn researchers with genetic data of large patient cohorts. “We were able to show that TP63 increases the activity of a number of genes that play a role in the development of isolated forms of cleft lip and palate,” said Ludwig.
Furthermore, TP63 changes the structure of the chromatin—the complex consisting of DNA and various proteins. Normally, the chromatin thread forms a compact bundle in the cell nucleus. However, when TP63 attaches to the thread, the ball loosens slightly in this area. Together with other modifications, this can lead to the increased reading of a specific gene in this region.
“According to current knowledge, TP63 regulates several thousand locations in our genome in this way,” said co-author Dr Julia Welzenbach from the University of Bonn. “Among them are 17 which, as we already know from large genetic studies, are involved in the formation of scissures and a large number of other regions whose involvement was previously unknown.”
The large number of regions activated by TP63 shows how important this gene is in humans. If its function is severely impaired by a mutation, this defect usually affects a large number of organs. This explains why TP63 was originally only associated with the syndromic form of cleft lip and palate. “In the case of the non-syndromic form, however, its disturbing activity is limited to developing facial cells,” said Ludwig.
The newly established cell system provides scientists with a tool with which they can investigate the biological causes of this developmental disorder in more detail. “For example, we could test the effect of different environmental factors on TP63 activity in facial cells,” stated Ludwig. It is known that environmental influences can increase the risk of facial malformation. Therefore, in the long term, such tests might lead to recommendations for better individual prophylaxis, applying in particular to families which have a genetic predisposition.
The study, titled “p63 establishes epithelial enhancers at critical craniofacial development genes”, was published in the May 2019 issue of Science Advances.
Tags:
Last month, Dental Tribune International (DTI) covered a new study on how lipid metabolism and maternal weight influence orofacial cleft (OFC) risk in ...
WASHINGTON, D.C., USA: As all phases of fetal development, palate growth is a marvel of nature. In one part of the process, ledges of tissue on the sides of...
MELBOURNE, Australia: In the first large-scale study to look at the oral microbiome, researchers from Murdoch Children’s Research Institute (MCRI) have ...
CHICAGO, US: Although manufacturers indicate that healing abutments (HAs) are single-use components, HAs are sometimes reused in clinical practice. Academic...
CLEVELAND, U.S.: Obesity and periodontal disease remain the most common noncommunicable diseases in the U.S. A recent study has explored the effect of ...
OKAYAMA, Japan: In a recent study, researchers from Okayama University investigated whether involuntary masseter muscle activity showed any specific pattern...
TÜBINGEN, Germany: Given that teeth are often the best-preserved part of human skeletons, researchers across many disciplines frequently rely on them to ...
KUOPIO, Finland: It is well established that coeliac disease can adversely affect oral health, yet the underlying mechanisms are not fully understood. A ...
PHILADELPHIA, Penn., US: A good oral hygiene routine requires manual dexterity and can be difficult for elderly people and people with disabilities. ...
CLEVELAND, U.S.: The negative health impacts of smoking are widely known; however, few have researched its consequences regarding endodontics. In a new ...
Live webinar
Wed. 3 June 2026
1:00 pm EST (New York)
Live webinar
Thu. 4 June 2026
2:00 pm EST (New York)
Live webinar
Mon. 8 June 2026
12:00 pm EST (New York)
Live webinar
Mon. 8 June 2026
1:00 pm EST (New York)
Dr. Anthony Mak B.D.S, Prof. Marleen Peumans
Live webinar
Mon. 8 June 2026
2:00 pm EST (New York)
Live webinar
Wed. 10 June 2026
11:00 am EST (New York)
Live webinar
Wed. 10 June 2026
2:00 pm EST (New York)
Nacho Fernández-Baca DDS, MSc



 Austria / Österreich
Austria / Österreich
 Bosnia and Herzegovina / Босна и Херцеговина
Bosnia and Herzegovina / Босна и Херцеговина
 Bulgaria / България
Bulgaria / България
 Croatia / Hrvatska
Croatia / Hrvatska
 Czech Republic & Slovakia / Česká republika & Slovensko
Czech Republic & Slovakia / Česká republika & Slovensko
 France / France
France / France
 Germany / Deutschland
Germany / Deutschland
 Greece / ΕΛΛΑΔΑ
Greece / ΕΛΛΑΔΑ
 Hungary / Hungary
Hungary / Hungary
 Italy / Italia
Italy / Italia
 Netherlands / Nederland
Netherlands / Nederland
 Nordic / Nordic
Nordic / Nordic
 Poland / Polska
Poland / Polska
 Portugal / Portugal
Portugal / Portugal
 Romania & Moldova / România & Moldova
Romania & Moldova / România & Moldova
 Slovenia / Slovenija
Slovenia / Slovenija
 Serbia & Montenegro / Србија и Црна Гора
Serbia & Montenegro / Србија и Црна Гора
 Spain / España
Spain / España
 Switzerland / Schweiz
Switzerland / Schweiz
 Turkey / Türkiye
Turkey / Türkiye
 UK & Ireland / UK & Ireland
UK & Ireland / UK & Ireland
 Brazil / Brasil
Brazil / Brasil
 Canada / Canada
Canada / Canada
 Latin America / Latinoamérica
Latin America / Latinoamérica
 USA / USA
USA / USA
 China / 中国
China / 中国
 India / भारत गणराज्य
India / भारत गणराज्य
 Pakistan / Pākistān
Pakistan / Pākistān
 Vietnam / Việt Nam
Vietnam / Việt Nam
 ASEAN / ASEAN
ASEAN / ASEAN
 Israel / מְדִינַת יִשְׂרָאֵל
Israel / מְדִינַת יִשְׂרָאֵל
 Algeria, Morocco & Tunisia / الجزائر والمغرب وتونس
Algeria, Morocco & Tunisia / الجزائر والمغرب وتونس
 Middle East / Middle East
Middle East / Middle East


































To post a reply please login or register